Possible new method identified to assess severity of TB infection
Metabolite of TB-causing bacteria also may help monitor response to treatment
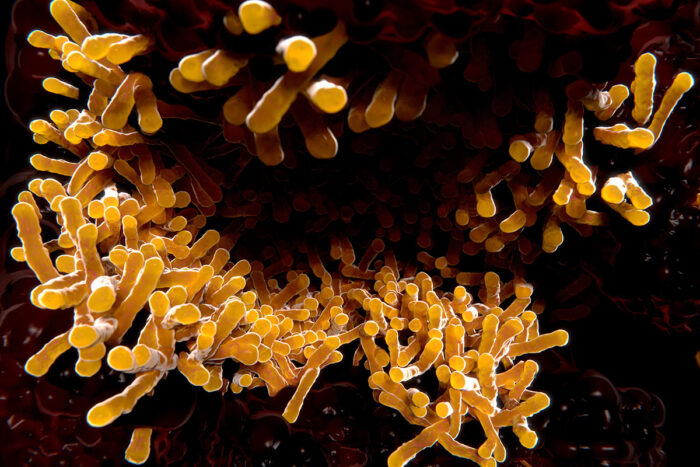 Getty Images
Getty ImagesPictured is an electron micrograph of Mycobacterium tuberculosis, the bacterium that causes tuberculosis. Researchers at Washington University School of Medicine in St. Louis have discovered a molecule in the lungs of tuberculosis (TB) patients that reflects the levels of TB-causing bacteria in the lungs. The findings may represent a step toward more tailored treatment regimens and better outcomes for TB patients.
Researchers at Washington University School of Medicine in St. Louis have discovered a molecule in the lungs of tuberculosis (TB) patients that reflects the levels of TB-causing bacteria in the lungs, information needed to better address the notoriously difficult to treat disease.
The findings, published Feb. 1 in the Journal of Clinical Investigation, could be a step toward more tailored treatment regimens and better outcomes for TB patients.
Most of the 10 million people who develop TB every year undergo months of treatment with multiple antibiotics, and despite the onerous drug regimen, roughly 5% of TB patients relapse within two years. Doctors don’t have a good way to track levels of bacteria in their lungs, so instead they monitor symptoms.
But the new research identifies a molecule in people with active TB and shows that levels of this molecule are higher in those with more severe infections. The molecule is a derivative of cholesterol known as cholestenone.
“It’s hard to guide clinical care and make treatment decisions when you don’t have a good way to measure whether you’re really reducing the amount of bacteria,” said senior author Jennifer A. Philips, MD, PhD, an associate professor of medicine and of molecular microbiology. “Lessened coughing, lower fevers, weight gain and overall feeling stronger are all good signs that what you are doing is working, but they are not proof that the bacteria have been eliminated. Some of the drugs that we use have a lot of toxicity, and we have to use multiple drugs for a long time. We’re probably overtreating a whole bunch of people, but we can’t distinguish the people who could stop taking them early from the people who need to go longer, or they’re going to relapse.”
Like all living things, Mycobacterium tuberculosis, the bacterium that causes TB, has to eat. It consumes chemical compounds and releases byproducts as it breaks the compounds down for energy, a process known as metabolism. Philips and colleagues — including co-first authors Pallavi Chandra, an instructor in medicine with a doctorate in microbiology, and Héloise Coullon, a researcher with doctorates in pharmacy and microbiology — reasoned that they might be able to identify infected cells and people by detecting the presence of byproducts of bacterial metabolism.
To search for such compounds, the researchers infected mouse cells with TB bacteria and then used mass spectrometry to identify metabolic compounds released over the next five days. Some cells were left uninfected for comparison. The researchers found more than 30 compounds that distinguished infected from uninfected cells. One of the most striking differences was in levels of cholestenone, a byproduct of bacterial cholesterol metabolism. TB bacteria can metabolize cholesterol, but they do not make it themselves; they acquire it from people during infection. People also metabolize cholesterol but through a different process that doesn’t generate cholestenone. Thus, cholestenone is a unique co-metabolite produced by bacteria during infection, making it a promising candidate to identify people with active cases of TB and measure the quantity of bacteria in their lungs.
The researchers studied 80 people, half in Vietnam and half in Peru, where TB is more common than in the U.S. All participants had symptoms of lung infection such as fevers, coughing and weight loss, but only half had TB. The remaining 40 people – 20 from each country – had other reasons for their symptoms, such as other kinds of lung infections.
With the help of Xuntian Jiang, PhD, an assistant professor of medicine and director of the Washington University Metabolomics Facility, the researchers analyzed sputum samples, produced deep in the lungs, from all 80 participants. The TB patients had significantly higher levels of cholestenone in their sputum than people without TB. And among the TB patients, those with more TB bacteria in their lungs had higher levels of cholestenone.
Sputum isn’t ideal for monitoring levels of TB bacteria in the lungs because acquiring it for analysis requires that the patient cough intensely to bring it up from deep in the lungs and then spit it out. Some people, especially children, simply can’t manage it. But blood tests are a routine part of medical care. If TB patients have high levels of cholestenone in the blood like they do in sputum, it might be possible to create a blood-based way to measure TB bacteria levels and monitor the effectiveness of treatment.
Philips and colleagues tested cholestenone levels in blood samples from all 80 participants. Of the people in Peru, the researchers were able to distinguish those with and without TB by their blood cholestenone levels. Of those in Vietnam, however, levels were not different between the two groups.
“It’s really exciting to find a difference in the levels in the patients in Peru,” Philips said. “We are really interested in understanding what is different about the two populations because then we might be able to find a different cholesterol metabolite to measure that is a blood biomarker of bacterial levels in all TB patients. There could be a lot of reasons why it worked in Peru but not Vietnam. It may be due to differences between people in Vietnam and Peru in terms of their genetics or the bacteria in their microbiomes. The TB bacteria themselves are of different lineages in Peru and in Vietnam, so that may be a factor, too. We’re continuing this project and exploring as many of these possibilities as we can.”