Bowman leading international supercomputing project
Crowd-sourced computer network delves into protein structure, seeks new disease therapies
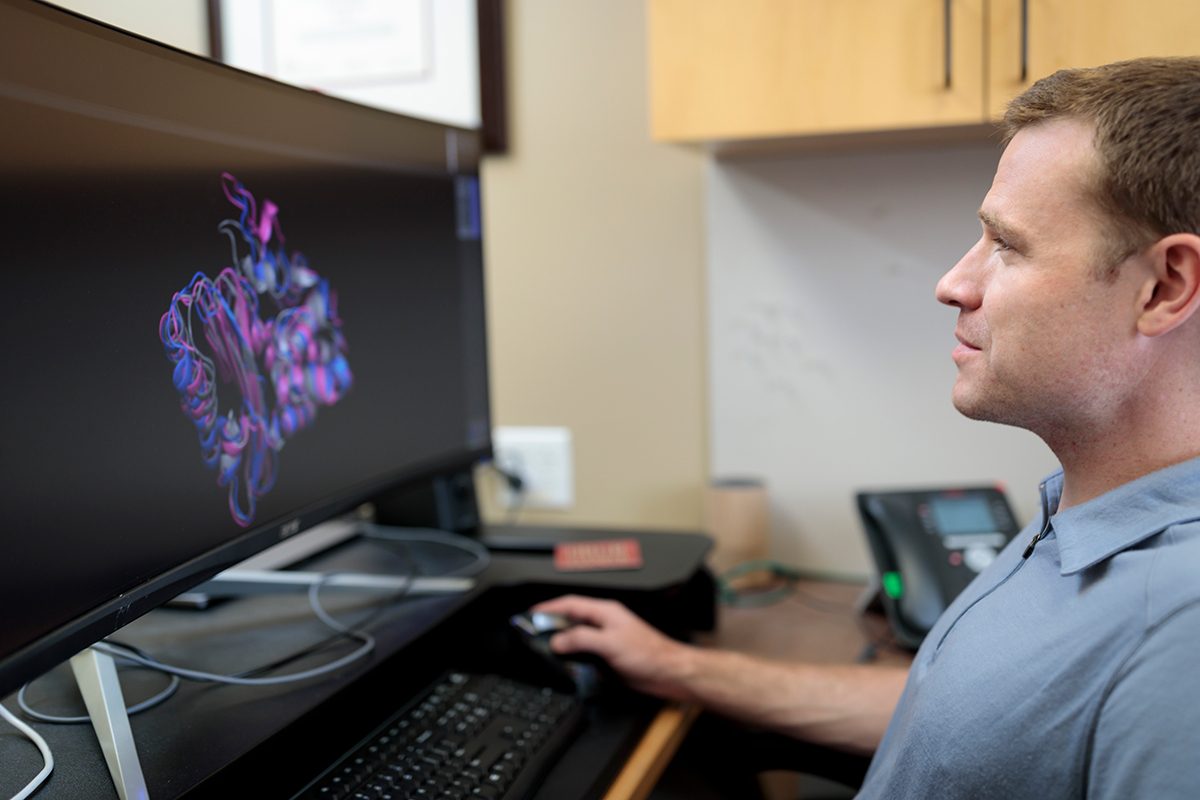
Huy Mach and Gaia RemerowskiGreg Bowman, PhD, an assistant professor of biochemistry and molecular biophysics at Washington University School of Medicine in St. Louis, is leading a supercomputing project called Folding@home. The project seeks to unravel the mysteries of protein dynamics, including the folding process, and their roles in health and disease.
When Greg Bowman presents a slideshow about the proteins he studies, their 3D shapes and folding patterns play out as animations on a big screen. As he describes these molecules, it might be easy to miss the fact that he can’t really see his own presentation, at least not the way the audience does.
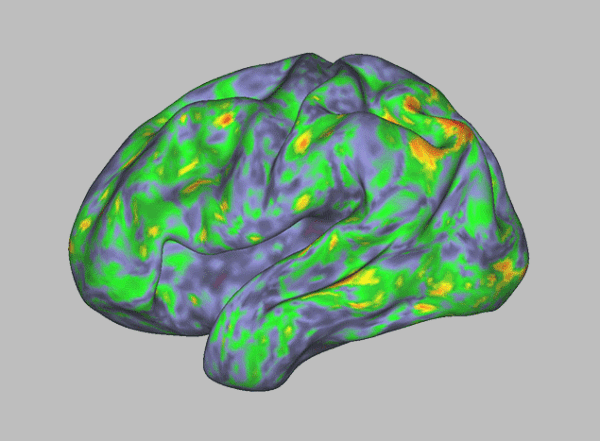
Bowman, PhD, an assistant professor of biochemistry and molecular biophysics at Washington University School of Medicine in St. Louis, is legally blind. He also now leads one of the largest crowd-sourced computational biology projects in the world. The effort is aimed at understanding how proteins fold into their proper shapes and the structural motions they undergo as they do their jobs keeping the body healthy. Proteins are vital cellular machinery, and understanding how they assemble and function — or malfunction — could shed light on many of the most vexing problems in medical science, from preventing Alzheimer’s disease, to treating cancer, to combating antibiotic resistance.
Appropriately called Folding@home, the project relies on the power of tens of thousands of home computers to perform the complex calculations required to simulate the dynamics of the proteins Bowman and his colleagues are studying. With this networked computing power, Folding@home is, essentially, one of the world’s largest supercomputers.
“There are some traditional supercomputing folks who might take issue with that characterization,” Bowman said with a laugh. “Rather than a single massive machine, Folding@home is a distributed computing network. Thousands of volunteers all over the world download our software and contribute a portion of their home computer setups to the project. But in terms of raw computing power — the sheer number of calculations it can perform per second — it’s on par with the world’s biggest supercomputers.”
Bowman got started on this work in the lab of Folding@home founder Vijay Pande, PhD, of Stanford University. Bowman earned his doctoral degree at Stanford and did postdoctoral research there. After 18 years at the helm, Pande chose Bowman to take over leadership, bringing Folding@home into the next decade and beyond.
 Matt Miller
Matt Miller“Greg has a unique combination of skills,” Pande said. “He has the technical chops to lead this complex project, and he has the people skills to manage the distributed nature of it, especially the fact that it involves so many different kinds of people — scientists and nonscientists alike. Greg also has great vision for the future of this project. He not only will keep the trains running on time, he has a strong picture of where Folding@home should be 10 to 20 years from now.”
Folding@home’s massive computing capacity is crucial to understanding protein folding, a problem Bowman calls a classic grand challenge in biochemistry and biophysics. Proteins are the raw materials that make up our bodies. But they are also the molecular machines that do the work of building those bodies and making sure they run properly. To do its work, a protein must fold into its proper form. If it doesn’t, things go wrong.
Bowman understands this more than most.
Born with normal vision, Bowman progressively lost sight, becoming legally blind by age 9 due to an inherited condition called Stargardt disease. A form of juvenile macular degeneration, it is caused when a protein that removes waste from cells in the retina doesn’t fold properly and can’t do its job. As a result, light-sensing cells in the retina become overwhelmed with waste and die, causing loss of central vision.
 Matt Miller
Matt MillerBowman said the experience lit a passion for biology and a drive to understand what goes wrong when the proteins our body relies on don’t work properly. Ultimately, he would like to find ways to fix them. But as a young student, Bowman quickly realized his route into the field might look a little different than that taken by the average biologist.
“I learned that experimental biology is not very accessible to people who are visually impaired,” Bowman said. “Essentially, I see at low resolution, mostly with my peripheral vision. I can navigate hallways and laboratories, but I can’t read the small dial on a pipette, for example.
“As I came to realize this, I also fell in love with computers,” he said. “I saw that the skills of computer science and mathematical modeling could be applied to biological problems. Plus, one of the many beauties of computers is that it’s really easy to zoom in on things. I can zoom in to 16 times magnification and scroll around on the screen, so I can read a scientific paper — or even just an email — for example.”
With Folding@home, Bowman and his colleagues are zooming in on proteins and how they fold much more than 16 times. Indeed, they are getting as close as physically possible — down to the atomic level. With this networked supercomputer, scientists can model proteins at the level of individual atoms in a fraction of the time it might take even powerful single computers. Many important biological processes that proteins perform take place over milliseconds to a few seconds. That might seem short, but measuring atoms as they bounce off one another requires time scales in femtoseconds — one quadrillionth of a second.
“To model just one millisecond of folding, even for an average-size protein, on a top-of-the-line MacBook Pro, it would take something like 500 years,” Bowman said. “But with Folding@home, we can split these problems into many independent chunks. We can send them to 1,000 people at the same time. Running those calculations in parallel, we can take these problems that would have taken 500 years and instead solve them in six months.”
As of this writing, Folding@home has more than 110,000 volunteer “folders” around the world who have shared a portion of their home computing capacity. According to videos from some volunteers, their reasons for contributing to the project are, like Bowman’s, personal. The program gives users some choice in what kinds of projects they contribute to, whether they are interested in boosting cancer research, preventing Alzheimer’s disease or fighting antibiotic resistance, among others.
 Matt Miller
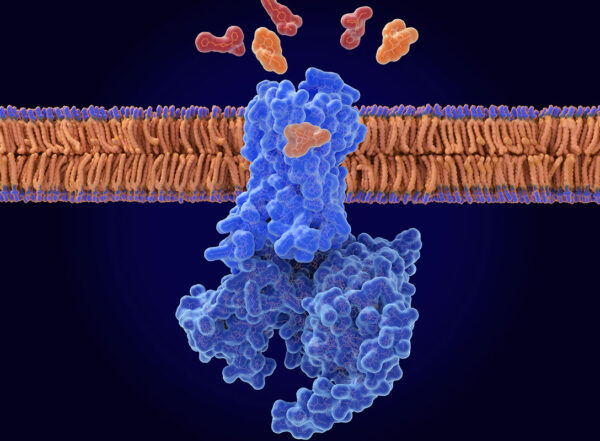
Matt MillerBowman envisions a future where Folding@home serves as a starting point for new drug design. Right now, scientists often have only one well-known protein structure to study. Beta lactamase, for example, is a protein that some bacteria deploy to protect themselves from antibiotics like penicillin. The protein has a well-documented, long-studied structure. But that structure only represents a single snapshot of beta lactamase at one moment in time.
“That snapshot contains valuable information,” Bowman said. “But it’s kind of like seeing a picture of a construction vehicle in a parking lot and trying to guess what it does. Really, what you would like is to watch this thing move around and see how it works together with other machinery to, say, build a building. We’re interested in watching how every atom in a protein moves — as it’s being assembled for the first time and as it goes about its jobs. The atoms in a protein are never still, they’re constantly jostling and moving around. And one genetic mutation changes maybe a dozen atoms out of thousands. We want to understand what that does to the entire protein.”
Among several projects, Bowman’s own lab is using Folding@home to seek new drugs to combat antibiotic resistance. Watching the movement of beta lactamase, for example, already has revealed what Bowman calls “cryptic pockets,” weak points in the protein that could be targeted by drugs but that are not visible in the long-studied snapshot of this protein. The cryptic pockets only reveal themselves when the protein is moving.
As Bowman sees the world a bit differently than most, Folding@home offers scientists a different look at long-studied proteins, revealing solutions to biological problems that might otherwise remain hidden from view.
To put your own computer to work folding proteins, visit the Folding@home website.
 Matt Miller
Matt Miller